View
 |
IMC has various display windows as follows.
- GenBank / EMBL viewer
- Sequence Viewer
- Annotation Window
- Multiple Genome Alignment Viewer (Reference Map)
- Circular Genome Viewer · Editor
- Plasmid Map Viewer · Editor
- Trace Viewer · Editor
- Phylogenetic Tree Viewer
- Alignment Viewer
- Label and Color Settings
|
Subcategories
- A viewer that can view the currently loaded sequence file in its original format.
- When you click on an sequence or annotation, the main feature map will display a map of that position.
- Conversely, with this viewer open, you can click on a part of the main feature map to display the text at the corresponding position in the file.
- A viewer that displays a sequence of the area currently displayed in the main feature map.
- You can display the display range on the viewer, show / hide the reverse strand, display / not display the amino acid translation, change the display method of the scale, display number of bases displayed.
- You can output image files in PDF, PNG, EMF format respectively.
| |
|
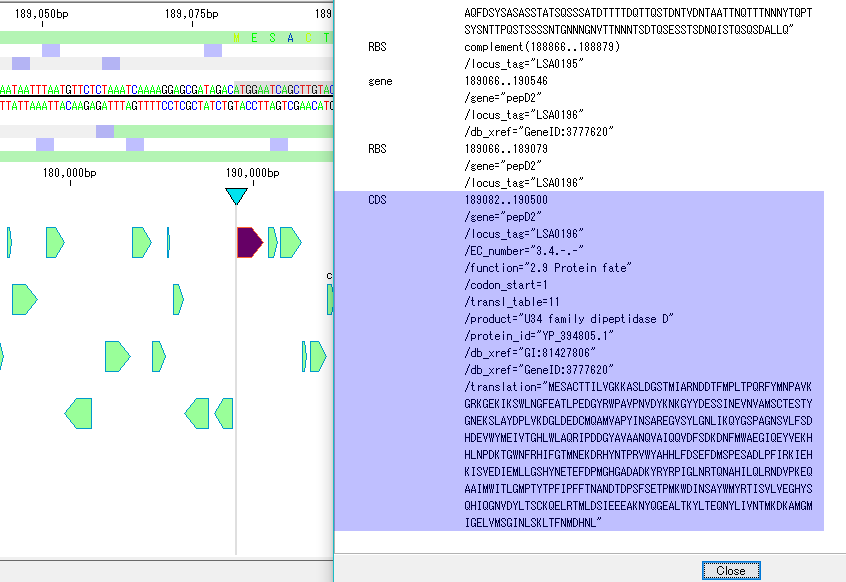
- A tool prepared for genome annotation. It is mainly used for annotating CDS.
|
| |
|
|
| |
|
- The window is divided into three parts, you can enter annotations directly, you can directly edit them, you can link the results of the homology search to the CDS features themselves and display them and post them.
- At the same time existing features can be selectively displayed in the sequence display area.
- You can change the display settings for sequences, features, and tick marks.
- Image contents can be output in PDF, PNG, EMF format.
- From this window you can also perform a homology search that queries this feature and the results are linked immediately.
|
- Editing / deleting of profile concentric circles, addition / editing / deletion of profile concentric circles, automatic map creation of multiple genomes, printing · image file output (PDF, PNG, EMF) is possible.
- The designed circular layout can be saved as a layout style and another genome sequence can be created with similar design.
- You can add concentric circles of various styles to the map.
- For one concentric circle, you can display and set multiple features, and you can specify strand designation, color classification, only specified features.
- Also, you can freely set the resolution such as content map such as GC content.
- It is also possible to draw by sequentially switching genomic sequences loaded in the current directory.
- You can change map resolution, diameter, phone size, and map name.
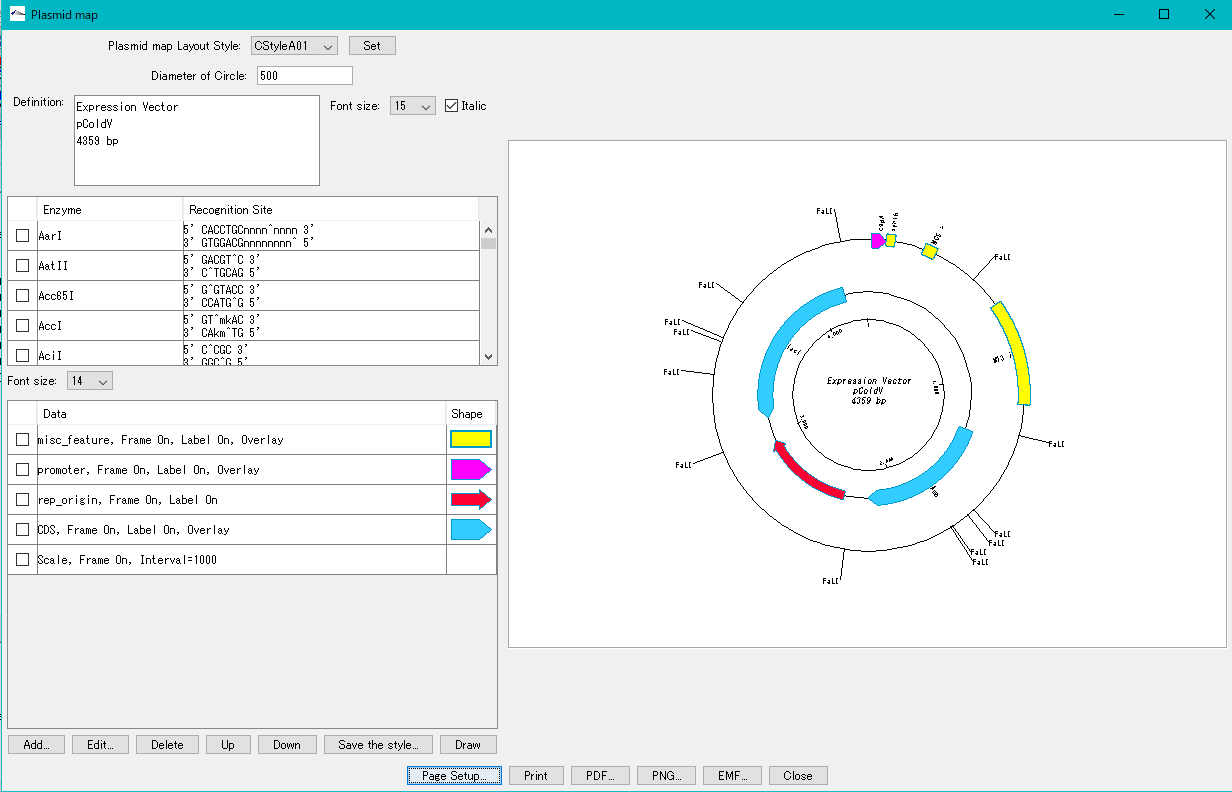
- It is similar to the circular genome map editor, but draws a relatively small circular genome.
- The annular lanes are arranged concentrically, and you can place multiple feature keys on one concentric circle.
- Strand, frame, feature shape, color designation of fill and frame, thickness of frame, selection of color classification, display / non-display of label, selection of Qualifier to be used as a label, font size, two concentric circle overlay functions ,
- You can change the scale interval.
- Designed styles can be saved and applied to other plasmids.
- You can change the order of concentric circles, add / delete, and edit.
- Plasmid maps can be output in three types of image formats: PDF, PNG, and EMF.
- It is a viewer / editor which is used when reading output file from ABI, SCF capillary sequencer.
- Not only the base sequence, but also its waveform (trace) can be displayed.
- You can change the base call.
The description window is a window for entering and editing values in the feature's Qualifier.
It is launched with a right click on the feature and you can edit all the Qualifiers for that feature.
You can also change the Position of the feature.
In addition, you can change the Feature Key to which the feature belongs.
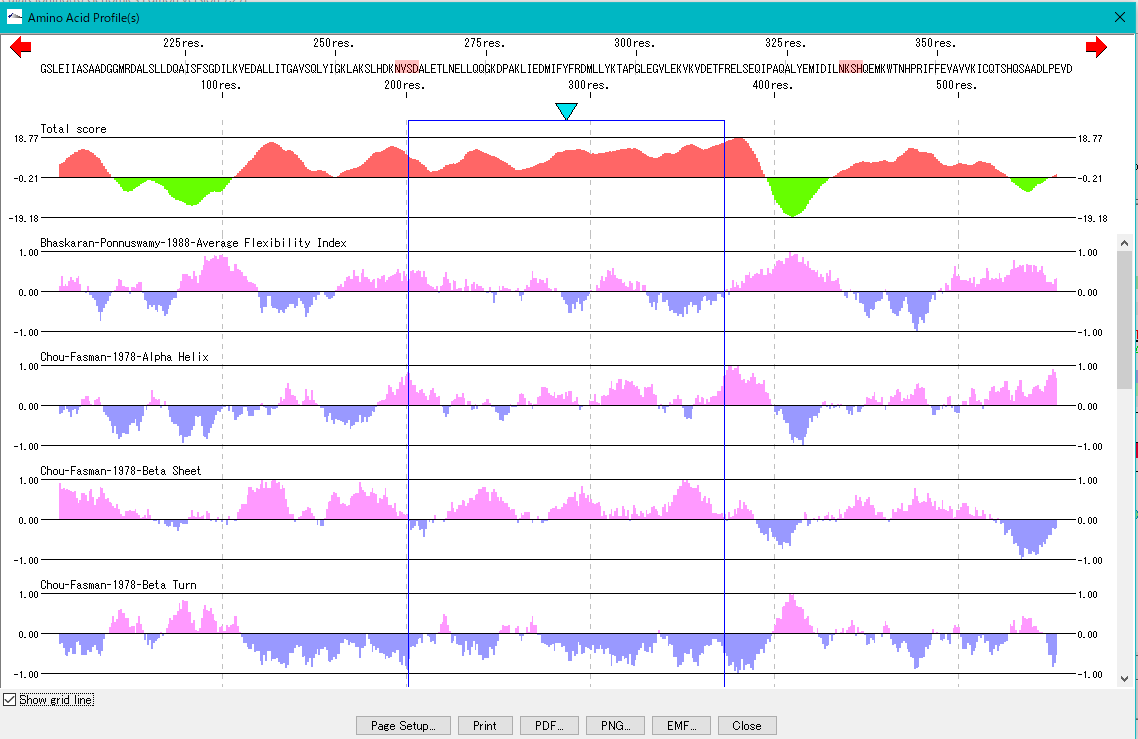
This is a viewer that displays profiles such as secondary structure of amino acid sequence in parallel. Total profile design and display, motif display and so on are possible.